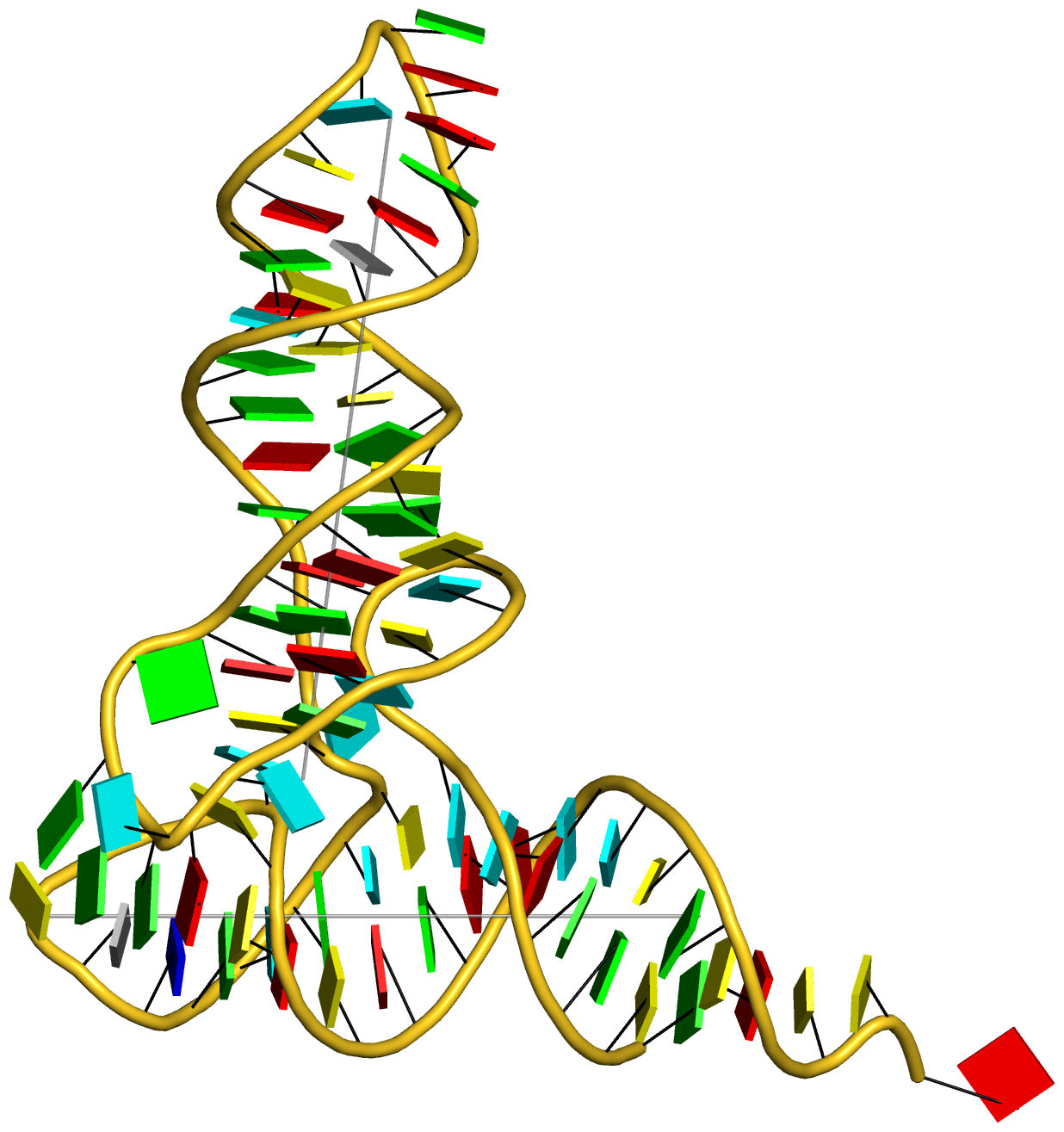
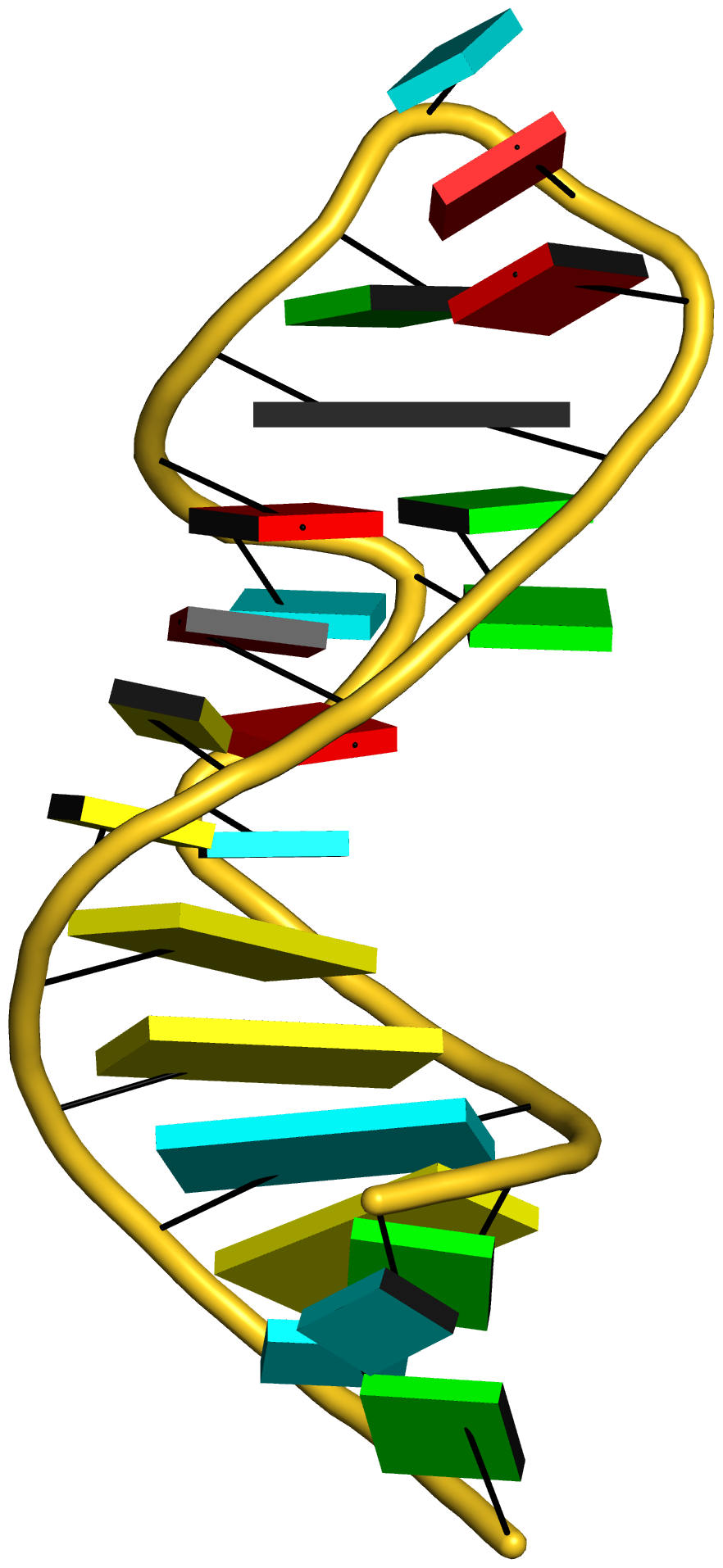
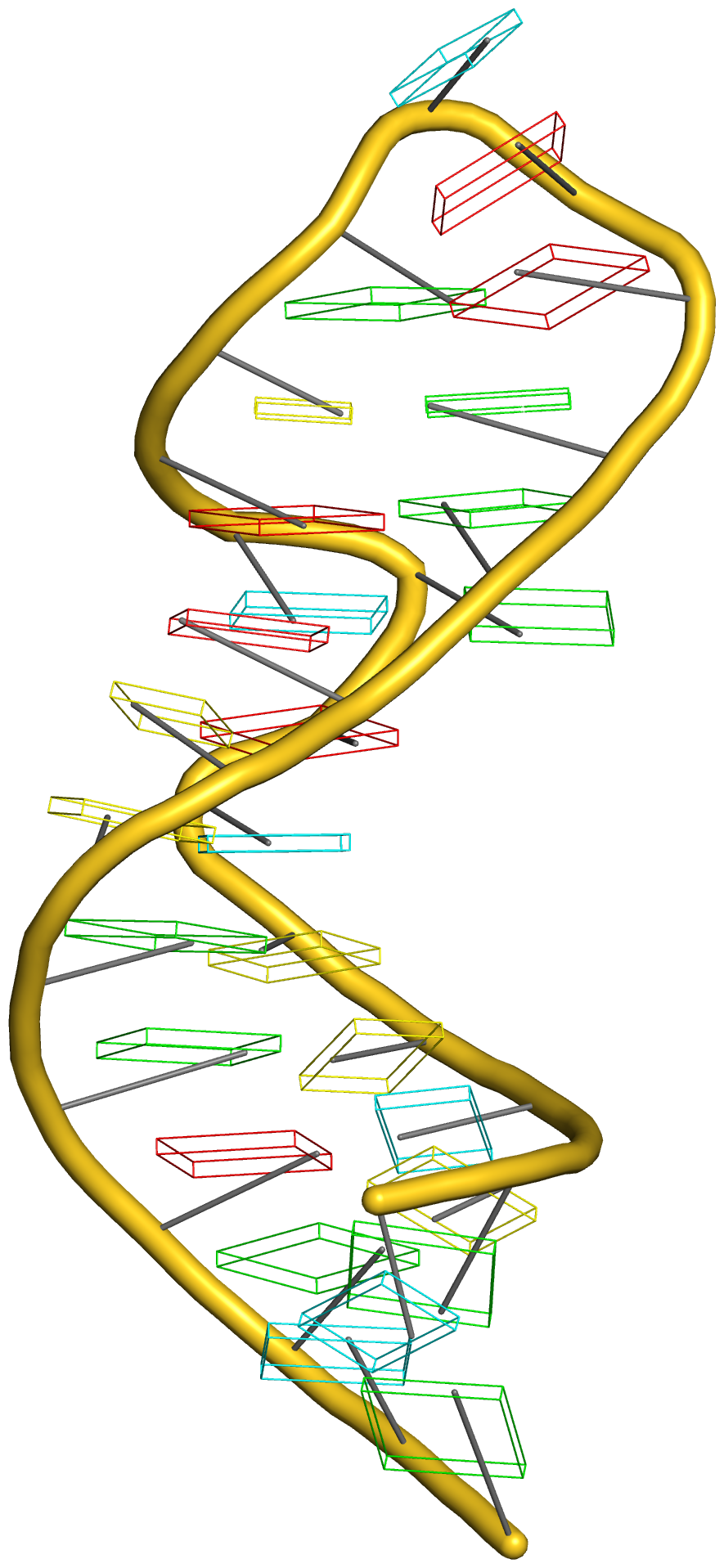
Crystal structure of SARS-CoV-2 stem–loop 5 (SL5) (PDB id: 9E9Q; Jones CP, Ferré-D'Amaré AR. 2025. Crystallographic and cryoEM analyses reveal SARS-CoV-2 SL5 is a mobile T-shaped four-way junction with deep pockets. RNA 31: 949–960). The T-shaped four-way junction of the coronavirus SL5 structural element provides a starting point for examining the structures of larger RNA motifs and their interactions with other molecules. Image highlighting the four arms of the junction. The RNA backbone is depicted by a gray ribbon. The bases within the arms of the junction are colored respectively in blue, red, yellow, and cyan. Cover image provided by X3DNA-DSSR, an NIGMS National Resource for Structural Bioinformatics of Nucleic Acids (R24GM153869; skmatics.x3dna.org). Image generated using DSSR and PyMOL (Lu XJ. 2020. Nucleic Acids Res 48: e74).
As the developer of DSSR, I am thrilled to see its application in cutting-edge research across multiple disciplines. Below is a list of four recent publications that highlight how DSSR has been utilized, underscoring its versatility and significance in structural bioinformatics.
In the Geng et al. (2025) Nucleic Acids Research (NAR) paper, titled 'Revealing hidden protonated conformational states in RNA dynamic ensembles', DSSR is simply cited as follows:
All bp geometries, hydrogen-bond, backbone, stacking, and sugar dihedral angles were calculated using X3DNA-DSSR [77].
In the preprint by Gordan et al. (2025), titled 'High-throughput characterization of transcription factors that modulate UV damage formation and repair at single-nucleotide resolution', DSSR is cited as follows:
Step base stacking, base pair shift, base pair slide, interbase angle, pseudorotation angle, and sugar puckering classifications of nucleobases were computed using X3DNA-DSSR (v2.5.0)75. Base stacking was defined as the overlapping polygon area in Å2 when projecting the dipyrimidine base ring atoms (excluding exocyclic atoms) into the mean base pair plane76. The sugar ring pseudorotation phase angle of each pyrimidine was also calculated using X3DNA-DSSR as described by Altona, C. & Sundaralingam, M.77 Interbase angle was defined as sqrt(propeller2+buckle2) per the X3DNA-DSSR documentation.
Figure 6: TF Binding Induces Structural Distortion Favorable to UV Dimerization is highly informative, particularly panel (a), which illustrates the ensemble of structural parameters that predispose dipyrimidines to cyclobutane pyrimidine dimers (CPD) or 6-4 pyrimidine-pyrimidones (6-4 PP) formation. DSSR is designed as an integrated software tool, offering a comprehensive suite of structural parameters not found in any other single tool I am aware of. Despite this, the innovative use of DSSR by Gordan et al. exceeds my expectations and demonstrates its versatility.
In the preprint by Kubaney et al. (2025) from the Baker group, titled 'RNA sequence design and protein-DNA specificity prediction with NA-MPNN', DSSR is cited as follows:
On the pseudoknot subset, we evaluate additional structure‐ and reactivity‐based metrics. DSSR v2.3.241 is used to extract the ground‐truth secondary structure from the native crystal structures. For each designed sequence, RibonanzaNet predicts 2A3 reactivity profiles, from which we compute predicted OpenKnot scores (see https://github.com/eternagame/OpenKnotScore)31 using the predicted reactivity together with the DSSR ground truth.
In a recent NSMB paper from the Baker group, titled 'Computational design of sequence-specific DNA-binding proteins', 3DNA is cited as follows:
RIF docking of scaffolds onto DNA targets (DBP design step 1) Structures of B-DNA for each target (Supplementary Table 2) were generated by (1) using the DNA portion of PDB 1BC8 (ref. 60), PDB 1YO5 (ref. 61), PDB 1L3L (ref. 51) or PDB 2O4A (ref. 62) or (2) using the software X3DNA63, followed by a constrained Rosetta relax of the DNA structure.
Please note that 3DNA has been replaced by DSSR. The functionality for constructing B-DNA models, previously provided by 3DNA, is now directly available in DSSR via its fiber and rebuild modules.
In the preprint by Si et al. (2025), titled 'End-to-End Single-Stranded DNA Sequence Design with All-Atom Structure Reconstruction', DSSR is cited as follows:
Since ViennaRNA and NUPACK require secondary structures as input, we used DSSR35 to extract secondary structures from the corresponding ssDNA three-dimensional structures.
The above use cases are merely a sample of how DSSR is utilized in the scientific literature. It is reasonable to state that DSSR has emerged as a de facto standard tool within the field of nucleic acid structural bioinformatics. Overall, DSSR is a mature, robust, and efficient software product that is actively developed and maintained. I am committed to making DSSR synonymous with quality and value. Its unmatched functionality, usability, and support save users significant time and effort compared to alternative solutions.
DSSR is available free of charge for academic users. Additionally, it has been integrated into other high-profile bioinformatics resources, including NAKB, PDB-redo, and N•ESPript.
References
- Geng A, Roy R, Ganser L, Li L, Al-Hashimi HM. Revealing hidden protonated conformational states in RNA dynamic ensembles. Nucleic Acids Research. 2025;53:gkaf1366. https://doi.org/10.1093/nar/gkaf1366.
- Gordan R, Wasserman H, Chi B, Bohm K, Duan M, Sahay H, et al. High-throughput characterization of transcription factors that modulate UV damage formation and repair at single-nucleotide resolution. 2025. https://doi.org/10.21203/rs.3.rs-8197218/v1.
- Kubaney A, Favor A, McHugh L, Mitra R, Pecoraro R, Dauparas J, et al. RNA sequence design and protein–DNA specificity prediction with NA-MPNN. 2025. https://doi.org/10.1101/2025.10.03.679414.
- Glasscock CJ, Pecoraro RJ, McHugh R, Doyle LA, Chen W, Boivin O, et al. Computational design of sequence-specific DNA-binding proteins. Nat Struct Mol Biol. 2025;32:2252–61. https://doi.org/10.1038/s41594-025-01669-4.
- Si Y, Xu Y, Chen L. End-to-end single-stranded DNA sequence design with all-atom structure reconstruction. 2025. https://doi.org/10.64898/2025.12.05.692525.
From early on, the x3dna.org domain and its related sub-domains (e.g., for the forum and the web-interface to DSSR) has been served via shared hosting. By and large, this simple arrangement has worked quite well. Over the years, though, I’ve gradually realized some of its inherent limitations. One is the limited resources available to the 3DNA-related websites. Another is the accessibility issue from countries like China.
To remedy such issues, I’ve recently moved the 3DNA Forum and the web-interface to DSSR to a dedicated web server at Columbia University. Moreover, a duplicate copy of the 3DNA homepage is made available via http://home.x3dna.org hosted at Columbia. The three new websites have been verified to be accessible directly from China.
These updates on x3dna.org not only ensure global accessibility to 3DNA/DSSR, but also allow for more web services to be made available.

As of v1.3.3-2015sep03, DSSR outputs the reference frame of any base or base-pair (bp). With an explicit list of such reference frames, one can better understand how the 3DNA/DSSR bp parameters are calculated. Moreover, third-party bioinformatics tools can take advantage of the frames for further exploration of nucleic acid structures, including visualization.
Let’s use the G1–C72 bp (detailed below) in the yeast phenylalanine tRNA (1ehz) as an example:
1 A.G1 A.C72 G-C WC 19-XIX cWW cW-W
The standard base reference frame for A.G1 is:
{
rsmd: 0.008,
origin: [53.757, 41.868, 52.93],
x_axis: [-0.259, -0.25, -0.933],
y_axis: [-0.543, 0.837, -0.073],
z_axis: [0.799, 0.488, -0.352]
}
And the one for A.C72 is:
{
rsmd: 0.006,
origin: [53.779, 42.132, 52.224],
x_axis: [-0.402, -0.311, -0.861],
y_axis: [0.451, -0.886, 0.109],
z_axis: [-0.797, -0.345, 0.497]
}
The G1–C72 bp reference frame is:
{
rsmd: null,
origin: [53.768, 42, 52.577],
x_axis: [-0.331, -0.283, -0.9],
y_axis: [-0.497, 0.863, -0.089],
z_axis: [0.802, 0.418, -0.427]
}
The beauty of the DSSR JSON output is that the above information can be extracted on the fly. For example, the following commands extract the above frames:
x3dna-dssr -i=1ehz.pdb --json | jq '.ntParams[] | select(.nt_id=="A.G1") | .frame'
x3dna-dssr -i=1ehz.pdb --json | jq '.ntParams[] | select(.nt_id=="A.C72") | .frame'
x3dna-dssr -i=1ehz.pdb --json --more | jq .pairs[0].frame
Note that in JSON, the array is 0-indexed, so the first bp (G1–C72) has an index of 0. In addition to jq, I also used underscore to pretty-print the frames.

Standard nitrogenous bases in DNA and RNA (A, C, G, T, and U) are aromatic compounds, each with a planar geometry. In the analyses of three-dimensional (3D) nucleic acid structures, the planar bases are normally taken as rigid bodies. The relative geometry of the two bases in base pair (bp) can then be rigorously quantified by six rigid-body parameters (see figure below). The three translations along the _x_-, _y_- and _z_-axes are termed Shear, Stretch, and Stagger, respectively. The three corresponding rotations are called Buckle, Propeller (twist), and Opening.

3DNA is unique with its coupled analyze and rebuild programs. The former calculates six bp parameters given 3D atomic coordinates (in PDB or PDBx/mmCIF format), while the later takes a set of such parameters to generate the corresponding structure. The rigor of the description can be easily verified in two equivalent ways: the close to zero root-mean-square deviation (RMSD) between the rebuilt structure and the original coordinates, after a least-squares superposition; or the identical six bp parameters when the rebuilt structure is analyzed.
As is often the case, a concrete example would make the point clear. Here I am using the reverse Hoogsteen (rHoogsteen) bp between U8 and A14 (see image below) in the yeast phenylalanine tRNA (1ehz) as an example. The PDB atomic coordinates of the U8–A14 rHoogsteen pair, excluding backbone atoms except for C1′, is stored in file 1ehz-U8-A14.pdb.

find_pair 1ehz-U8-A14.pdb stdout | analyze stdin
# bp parameters in file '1ehz-U8-A14.out'
# also generated 'bp_step.par' for rebuilding below
rebuild -atomic bp_step.par 1ehz-U8-A14-3DNA.pdb
# rmsd is 0.044 Å between '1ehz-U8-A14.pdb' and '1ehz-U8-A14-3DNA.pdb'
find_pair 1ehz-U8-A14-3DNA.pdb stdout | analyze stdin
# bp parameters of the rebuilt structure in '1ehz-U8-A14-3DNA.out'
rebuild -atomic bp_step.par 1ehz-U8-A14-3DNA-new.pdb
# rmsd is 0 Å between '1ehz-U8-A14-3DNA.pdb' and '1ehz-U8-A14-3DNA-new.pdb'
Note that the above commands should be performed in order, since the file bp_step.par is overwritten after each analyze run. For your verification, here are the links to the five files:
The 0.044 Å rmsd between the original PDB coordinates in 1ehz-U8-A14.pdb and the 3DNA rebuilt structure in 1ehz-U8-A14-3DNA.pdb is due to the slight non-planarity of experimental bases. The rmsd is 0 between the two rounds of 3DNA rebuilt structures, 1ehz-U8-A14-3DNA.pdb and 1ehz-U8-A14-3DNA-new.pdb, as expected.
The bp parameters in 1ehz-U8-A14.out and 1ehz-U8-A14-3DNA.out are identical, as expected, and they are shown below.
Local base-pair parameters
bp Shear Stretch Stagger Buckle Propeller Opening
1 U-A 4.14 -1.91 0.77 -4.62 12.12 -103.09
Running DSSR on 1ehz-U8-A14.pdb gives the following results. Note that the six bp parameters (last row prefixed with bp-pars) are the exactly same as in 3DNA — we are consistent.
# x3dna-dssr -i=1ehz-U8-A14.pdb --more
List of 1 base pair
nt1 nt2 bp name Saenger LW DSSR
1 A.U8 A.A14 U-A rHoogsteen 24-XXIV tWH tW-M
[n/a(n/a) ---- lambda=28.3] [n/a(n/a) ---- lambda=21.5]
d(C1'-C1')=9.63 d(N1-N9)=7.06 d(C6-C8)=6.00 tor(C1'-N1-N9-C1')=174.4
H-bonds[2]: "O2(carbonyl)-N6(amino)[3.00],N3(imino)-N7[2.74]"
interBase-angle=12.97 Simple-bpParams: Shear=4.28 Stretch=1.55 Buckle=-11.8 Propeller=5.4
bp-pars: [4.14 -1.91 0.77 -4.62 12.12 -103.09]
As mentioned in the recent DSSR paper:
As in 3DNA (6,7), DSSR takes advantage of the six standard base-pair parameters––three translations (Shear, Stretch, Stagger) and three rotations (Buckle, Propeller, Opening)––to quantify the relative spatial position and orientation of any two interacting bases rigorously. Among the six parameters, only Shear, Stretch, and Opening are critical for characterizing different types of pairs. Buckle, Propeller and Stagger, on the other hand, describe the nonplanarity of a given pair (6). By virtue of the definition of the standard base reference frame, Shear, Stretch, and Opening are all close to zero for Watson-Crick pairs. Moreover, every other type of pair has a set of characteristic parameters. For example, the wobble G–U pair is characterized by an average Shear of –2.2 Å, and the Hoogsteen A+U pair is distinguished by a Stretch of approximately –3.5 Å and an Opening of near 66º.
In a follow-up post, I will talk about the “simple” bp parameters (Simple-bpParams in the above DSSR output list) recently introduced into DSSR — stay tuned!

As of DSSR v1.3.0-2015aug27, the --json option is available for producing analysis results that is strictly compliant with the JSON data exchange format. The JSON file contains numerous DSSR-derived structural features, including those in the default main output, backbone torsions in dssr-torsions.txt, and a detailed list of hydrogen bonds.
According to the official JSON website:
JSON (JavaScript Object Notation) is a lightweight data-interchange format. It is easy for humans to read and write. It is easy for machines to parse and generate. It is based on a subset of the JavaScript Programming Language… JSON is a text format that is completely language independent… These properties make JSON an ideal data-interchange language.
Indeed, the JSON output file makes DSSR readily accessible for integration with other bioinformatics tools or normal usages from the command line. Using the classic yeast phenylalanine tRNA 1ehz as an example (1ehz.pdb), let’s go over some simple use-cases. Note that the following examples take advantage of jq, a lightweight and flexible command-line JSON processor.
x3dna-dssr -i=1ehz.pdb --json -o=1ehz-dssr.json
jq . 1ehz-dssr.json # reformatted for pretty output
x3dna-dssr -i=1ehz.pdb --json | jq . # the above 2 steps combined
With 1ehz-dssr.json in hand, we can easily extract DSSR-derived structural features of interest:
jq .pairs 1ehz-dssr.json # list of 34 pairs
jq .multiplets 1ehz-dssr.json # list of 4 base triplets
jq .hbonds 1ehz-dssr.json # list of hydrogen bonds
jq .helices 1ehz-dssr.json
jq .stems 1ehz-dssr.json
# list of nucleotide parameters, including torsion angles and suites
jq .ntParams 1ehz-dssr.json
# list of 14 modified nucleotides
jq '.ntParams[] | select(.is_modified)' 1ehz-dssr.json
# select nucleotide id, delta torsion, sugar puckering and cluster of suite name
jq '.ntParams[] | {nt_id, delta, puckering, cluster}' 1ehz-dssr.json
# same selection as above, but in 'Comma Separated Values' format
jq -r '.ntParams[] | [.nt_id, .delta, .puckering, .cluster] | @csv' 1ehz-dssr.json
Here is the result of running jq (v1.5) to select multiplets:
# jq .multiplets 1ehz-dssr.json
[
{
"index": 1,
"num_nts": 3,
"nts_short": "UAA",
"nts_long": "A.U8,A.A14,A.A21"
},
{
"index": 2,
"num_nts": 3,
"nts_short": "AUA",
"nts_long": "A.A9,A.U12,A.A23"
},
{
"index": 3,
"num_nts": 3,
"nts_short": "gCG",
"nts_long": "A.2MG10,A.C25,A.G45"
},
{
"index": 4,
"num_nts": 3,
"nts_short": "CGg",
"nts_long": "A.C13,A.G22,A.7MG46"
}
]
With the JSON file, DSSR can now be connected with the bioinformatics community in a ‘structured’ way, with a clearly delineated boundary. Now I can enjoy the freedom of refining the default main output format, without worrying too much about breaking third-party parsers. Moreover, I no longer need to write an adapter for each integration of DSSR with other tools. So nice!
For your reference, here is the output file 1ehz-dssr.json. It may be possible that the identifiers (names) of the JSON output will be refined in the next few iterations. I welcome your comments to make the DSSR-derived JSON better suite your needs.

It is a great pleasure to note that a paper titled DSSR, an integrated software tool for dissecting the spatial structure of RNA has recently been published in Nucleic Acids Research (NAR). Co-authored by Harmen Bussemaker, Wilma Olson and me (a team with a unique combination of complementary expertise), this DSSR paper represents another solid piece of work that I feel proud of. In contrast to our previous GpU dinucleotide platform paper focusing on results, and the two major 3DNA papers concentrating on methods, the current NAR article describes significant scientific findings that are enabled by the novel analysis algorithms implemented in the program. Moreover, DSSR introduces an appealing and highly informative “cartoon-block” representation of RNA structures that combines PyMOL cartoon schematics with 3DNA base color-coded rectangular blocks.
The abstract of the paper is quoted below:
Insight into the three-dimensional architecture of RNA is essential for understanding its cellular functions. However, even the classic transfer RNA structure contains features that are overlooked by existing bioinformatics tools. Here we present DSSR (Dissecting the Spatial Structure of RNA), an integrated and automated tool for analyzing and annotating RNA tertiary structures. The software identifies canonical and noncanonical base pairs, including those with modified nucleotides, in any tautomeric or protonation state. DSSR detects higher-order coplanar base associations, termed multiplets. It finds arrays of stacked pairs, classifies them by base-pair identity and backbone connectivity, and distinguishes a stem of covalently connected canonical pairs from a helix of stacked pairs of arbitrary type/linkage. DSSR identifies coaxial stacking of multiple stems within a single helix and lists isolated canonical pairs that lie outside of a stem. The program characterizes ‘closed’ loops of various types (hairpin, bulge, internal, and junction loops) and pseudoknots of arbitrary complexity. Notably, DSSR employs isolated pairs and the ends of stems, whether pseudoknotted or not, to define junction loops. This new, inclusive definition provides a novel perspective on the spatial organization of RNA. Tests on all nucleic acid structures in the Protein Data Bank confirm the efficiency and robustness of the software, and applications to representative RNA molecules illustrate its unique features. DSSR and related materials are freely available at http://home.x3dna.org/.
During the review process, we are delighted that the referees confirmed the claim that we made in the cover letter: “We would also like to emphasize that our reported results are easily verifiable, and we assure rigorous reproducibility of the data and figures described in this article.” Now that the paper has been published, as a follow-up, I’ve made available all the scripts and data files associated with the paper in a new section DSSR-NAR paper on the 3DNA Forum. The DSSR User Manual has also been updated with additional, previously undocumented, auxiliary options.
Overall, it took me more than ten days to create the 19 posts in the DSSR-NAR paper section and to revise the DSSR User Manual, along with other minor refinements for consistency. During the process, I’ve tried to make the scripts and data files self-contained for wide accessibility and easy understanding.
Any interested party should now be able to reproduce the table and figures (including the supplementary data) reported in the article. Moreover, with the additional details given in the post RNA cartoon-block representations with PyMOL and DSSR, one can easily generate similar schematic images as shown below:
I feel confident to claim that the results reported in our DSSR paper are reproducible. If you have issues related to the paper, please post them on the 3DNA Forum. I strive to respond promptly to any questions asked there.
In summary, DSSR is an integrated computational tool, designed from the bottom up to streamline the analysis of RNA three-dimensional structures. It is built upon my extensive experience in supporting 3DNA, growing knowledge of RNA structures, and refined programming skills. DSSR has a combined set of functionalities well beyond the scope of any known specialized resources. The program may well serve as a cornerstone for RNA structural bioinformatics and will benefit a broad range of possible applications.

The conformation of the five-membered sugar ring in DNA/RNA structures can be characterized using the five corresponding endocyclic torsion angles (shown below).
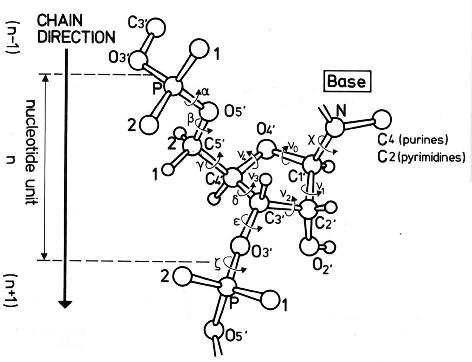
v0: C4'-O4'-C1'-C2'
v1: O4'-C1'-C2'-C3'
v2: C1'-C2'-C3'-C4'
v3: C2'-C3'-C4'-O4'
v4: C3'-C4'-O4'-C1'
On account of the five-member ring constraint, the conformation can be characterized approximately by 5 - 3 = 2 parameters. Using the concept of pseudorotation of the sugar ring, the two parameters are the amplitude (τm) and phase angle (P, in the range of 0° to 360°).
One set of widely used formula to convert the five torsion angles to the pseudorotation parameters is due to Altona & Sundaralingam (1972): “Conformational Analysis of the Sugar Ring in Nucleosides and Nucleotides. A New Description Using the Concept of Pseudorotation” [J. Am. Chem. Soc., 94(23), pp 8205–8212]. As always, the concept is best illustrated with an example. Here I use the sugar ring of G4 (chain A) of the Dickerson-Drew dodecamer (1bna), with Matlab/Octave code:
# xyz coordinates of the sugar ring: G4 (chain A), 1bna
ATOM 63 C4' DG A 4 21.393 16.960 18.505 1.00 53.00
ATOM 64 O4' DG A 4 20.353 17.952 18.496 1.00 38.79
ATOM 65 C3' DG A 4 21.264 16.229 17.176 1.00 56.72
ATOM 67 C2' DG A 4 20.793 17.368 16.288 1.00 40.81
ATOM 68 C1' DG A 4 19.716 17.901 17.218 1.00 30.52
# endocyclic torsion angles:
v0 = -26.7; v1 = 46.3; v2 = -47.1; v3 = 33.4; v4 = -4.4
Pconst = sin(pi/5) + sin(pi/2.5) # 1.5388
P0 = atan2(v4 + v1 - v3 - v0, 2.0 * v2 * Pconst); # 2.9034
tm = v2 / cos(P0); # amplitude: 48.469
P = 180/pi * P0; # phase angle: 166.35 [P + 360 if P0 < 0]
The Altona & Sundaralingam (1972) pseudorotation parameters are what have been adopted in 3DNA, following the NewHelix program of Dr. Dickerson. The Curves+ program, on the other hand, uses another (newer) set of formula due to Westhof & Sundaralingam (1983): “A Method for the Analysis of Puckering Disorder in Five-Membered Rings: The Relative Mobilities of Furanose and Proline Rings and Their Effects on Polynucleotide and Polypeptide Backbone Flexibility” [J. Am. Chem. Soc., 105(4), pp 970–976]. The two sets of formula, by Altona & Sundaralingam (1972) and Westhof & Sundaralingam (1983), give slightly different numerical values for the two pseudorotation parameters (τm and P).
Since 3DNA and Curves+ are currently two of the most widely used programs for conformational analysis of nucleic acid structures, the subtle differences in pseudorotation parameters may cause confusions for users who use (or are familiar with) both programs. Over the past few years, I have indeed received such questions via email.
With the same G4 (chain A, 1bna) sugar ring, here is the Matlab/Octave script showing how Curve+ calculates the pseudorotation parameters:
# xyz coordinates of sugar ring G4 (chain A, 1bna)
# endocyclic torsion angles, same as above
v0 = -26.7; v1 = 46.3; v2 = -47.1; v3 = 33.4; v4 = -4.4
v = [v2, v3, v4, v0, v1]; # reorder them into vector v[]
A = 0; B = 0;
for i = 1:5
t = 0.8 * pi * (i - 1);
A += v(i) * cos(t);
B += v(i) * sin(t);
end
A *= 0.4; # -48.476
B *= -0.4; # 11.516
tm = sqrt(A * A + B * B); # 49.825
c = A/tm; s = B/tm;
P = atan2(s, c) * 180 / pi; # 166.64
For this specific example, i.e., the sugar ring of G4 (chain A, 1bna), the pseudorotation parameters as calculated by 3DNA per Altona & Sundaralingam (1972) and Curves+ per Westhof & Sundaralingam (1983) are as follows:
amplitude phase angle
3DNA 48.469 166.35
Curves+ 49.825 166.64
Needless to say, the differences are subtle, and few people will notice/bother at all. For those who do care about such little details, however, hopefully this post will help you understand where the differences actually come from.
For consistency with the 3DNA output, DSSR (by default) also follows the Altona & Sundaralingam (1972) definitions of sugar pseudorotation. Nevertheless, DSSR also contains an undocumented option, --sugar-pucker=westhof83, to output τm and P according to the Westhof & Sundaralingam (1983) definitions.
Each sugar is assigned into one of the following ten puckering modes, by dividing the phase angle (P, in the range of 0° to 360°) into 36° ranges reach.
C3'-endo, C4'-exo, O4'-endo, C1'-exo, C2'-endo,
C3'-exo, C4'-endo, O4'-exo, C1'-endo, C2'-exo
For sugars in nucleic acid structures, C3’-endo [0°, 36°) and C2’-endo [144°, 180°) are predominant. The former corresponds to sugars in ‘canonical’ RNA or A-form DNA, and the latter in sugars of standard B-form DNA. In reality, RNA structures as deposited in the PDB could also contain C2′-endo sugars. One significant example is the GpU dinucleotide platforms, where the 5′-ribose sugar (G) is in the C2′-endo form and the 3′-sugar (U) in the C3′-endo form — see my blog post, titled ‘Is the O2′(G)…O2P H-bond in GpU platforms real?’.
Notes:
- This post is based on my 2011-06-11 blog post with the same title.
- While visiting Lyon in July 2014, I had the opportunity to hear Dr. Lavery’s opinion on adopting the Westhof & Sundaralingam (1983) sugar-pucker definitions in Curves+. I learned that the new formula are more robust in rare, extreme cases of sugar conformation than the 1972 variants. After all, Dr. Sundaralingam is a co-author on both papers. It is possible that in future releases of DSSR, the new 1983 formula for sugar pucker would become the default.

In the DSSR v1.2.7-2015jun09 release, I documented two additional command-line options (--prefix and --cleanup) that are related to the various auxiliary files. As a matter of fact, these two options (among quite a few others) have been there for a long time, but without being explicitly described. The point is not to hide but to simplify — one of the design goals of DSSR is simplicity. DSSR has already possessed numerous key functionality to be appreciated. Before DSSR is firmly established in the RNA bioinformatics field, I beleive too many nonessential “features” could be distracting. While writing and refining the DSSR code, I do feel that some ‘auxiliary’ features could be handy for experienced users (including myself). So along the way, I’ve added many ‘hidden’ options that are either experimental or potentially useful.
On one side, I sense it is acceptable for a scientific software to actually does more than it claims. On the other hand, I have always been quick in addressing users’ requests — as one example, check for the --select option recently introduced into DSSR in response to a user request, and the ‘hidden’ --dbn-break option for specifying the symbol to separate multiple chains or chain breaks in DSSR-derived dot-bracket notation.
Back to --prefix and --cleanup, the purposes of these two closely related options can be best illustrated using the yeast phenylalanine tRNA structure (1ehz) as an example. By default, running x3dna-dssr -i=1ehz.pdb will produce a total of 11 auxiliary files, with names prefixed with dssr-, as shown below:
List of 11 additional files
1 dssr-stems.pdb -- an ensemble of stems
2 dssr-helices.pdb -- an ensemble of helices (coaxial stacking)
3 dssr-pairs.pdb -- an ensemble of base pairs
4 dssr-multiplets.pdb -- an ensemble of multiplets
5 dssr-hairpins.pdb -- an ensemble of hairpin loops
6 dssr-junctions.pdb -- an ensemble of junctions (multi-branch)
7 dssr-2ndstrs.bpseq -- secondary structure in bpseq format
8 dssr-2ndstrs.ct -- secondary structure in connect table format
9 dssr-2ndstrs.dbn -- secondary structure in dot-bracket notation
10 dssr-torsions.txt -- backbone torsion angles and suite names
11 dssr-stacks.pdb -- an ensemble of stacks
With ‘fixed’ generic names by default, users can run DSSR in a directory repeatedly without creating too many files. This practice follows that used in the 3DNA suite of programs. However, my experience in supporting 3DNA over the years has shown that users (myself included) may want to explore further some of the files, e.g. ‘dssr-multiplets.pdb’ for displaying the base multiplets (four triplets here). One could easily use command-line (script) to change a generic name to a more appropriate one: e.g., mv dssr-multiplets.pdb 1ehz-multiplets.pdb for 1ehz. A better solution, however, is by introducing a customized prefix to the additional files, and that’s exactly where the --prefix option comes in. The option is specified like this: --prefix=text where text can be any string as appropriate. So running x3dna-dssr -i=1ehz.pdb --prefix=1ehz, for example, will lead to the following output:
List of 11 additional files
1 1ehz-stems.pdb -- an ensemble of stems
2 1ehz-helices.pdb -- an ensemble of helices (coaxial stacking)
3 1ehz-pairs.pdb -- an ensemble of base pairs
4 1ehz-multiplets.pdb -- an ensemble of multiplets
5 1ehz-hairpins.pdb -- an ensemble of hairpin loops
6 1ehz-junctions.pdb -- an ensemble of junctions (multi-branch)
7 1ehz-2ndstrs.bpseq -- secondary structure in bpseq format
8 1ehz-2ndstrs.ct -- secondary structure in connect table format
9 1ehz-2ndstrs.dbn -- secondary structure in dot-bracket notation
10 1ehz-torsions.txt -- backbone torsion angles and suite names
11 1ehz-stacks.pdb -- an ensemble of stacks
The --cleanup option, as its name implies, is to tidy up a directory by removing the auxiliary files generated by DSSR. The usage is very simple:
x3dna-dssr --cleanup
x3dna-dssr --cleanup --prefix=1ehz
The former gets rid of the default ‘fixed’ generic auxiliary files (dssr-pairs.pdb etc), whilst the latter deletes prefixed supporting files (1ehz-pairs.pdb etc).

Recently, I came across and have been surprised by the different assignment of HETATM vs. ATOM records for modified nucleotides in PDB vs. PDBx/mmCIF format. As always, the issue is best illustrated with a concrete example. Here is what I observed in the PDB entry 1ehz, the crystal structure of yeast phenylalanine tRNA at 1.93 Å resolution.
DSSR identifies 14 modified nucleotides (of 11 types) in 1ehz as shown below:
List of 11 types of 14 modified nucleotides
nt count list
1 1MA-a 1 A.1MA58
2 2MG-g 1 A.2MG10
3 5MC-c 2 A.5MC40,A.5MC49
4 5MU-t 1 A.5MU54
5 7MG-g 1 A.7MG46
6 H2U-u 2 A.H2U16,A.H2U17
7 M2G-g 1 A.M2G26
8 OMC-c 1 A.OMC32
9 OMG-g 1 A.OMG34
10 PSU-P 2 A.PSU39,A.PSU55
11 YYG-g 1 A.YYG37
In file 1ehz.pdb downloaded from RCSB PDB, all the 14 modified nucleotides are assigned as HETATM whereas in 1ehz.cif the corresponding records are ATOM. Here is the excerpt for 1MA58 in PDB format:
HETATM 1252 P 1MA A 58 73.770 67.765 34.057 1.00 30.65 P
HETATM 1253 OP1 1MA A 58 72.638 67.886 33.105 1.00 32.84 O
HETATM 1254 OP2 1MA A 58 73.621 68.229 35.450 1.00 29.49 O
HETATM 1255 O5' 1MA A 58 74.315 66.273 34.254 1.00 28.81 O
HETATM 1256 C5' 1MA A 58 74.592 65.439 33.080 1.00 29.42 C
HETATM 1257 C4' 1MA A 58 74.279 63.972 33.383 1.00 33.42 C
HETATM 1258 O4' 1MA A 58 74.880 63.685 34.667 1.00 32.36 O
HETATM 1259 C3' 1MA A 58 72.789 63.573 33.509 1.00 35.13 C
HETATM 1260 O3' 1MA A 58 72.625 62.168 33.250 1.00 36.80 O
HETATM 1261 C2' 1MA A 58 72.560 63.667 35.012 1.00 34.80 C
HETATM 1262 O2' 1MA A 58 71.525 62.828 35.506 1.00 36.27 O
HETATM 1263 C1' 1MA A 58 73.908 63.150 35.551 1.00 33.62 C
HETATM 1264 N9 1MA A 58 74.284 63.494 36.930 1.00 30.36 N
HETATM 1265 C8 1MA A 58 73.887 64.574 37.688 1.00 34.55 C
HETATM 1266 N7 1MA A 58 74.415 64.610 38.899 1.00 33.32 N
HETATM 1267 C5 1MA A 58 75.204 63.469 38.953 1.00 33.37 C
HETATM 1268 C6 1MA A 58 76.031 62.941 39.948 1.00 33.58 C
HETATM 1269 N6 1MA A 58 76.184 63.488 41.134 1.00 41.19 N
HETATM 1270 N1 1MA A 58 76.708 61.803 39.669 1.00 34.48 N
HETATM 1271 CM1 1MA A 58 77.649 61.222 40.626 1.00 31.43 C
HETATM 1272 C2 1MA A 58 76.527 61.216 38.479 1.00 28.43 C
HETATM 1273 N3 1MA A 58 75.793 61.624 37.453 1.00 31.67 N
HETATM 1274 C4 1MA A 58 75.142 62.771 37.747 1.00 33.02 C
The corresponding section in PDBx/mmCIF format is:
ATOM 1252 P P . 1MA A 1 58 ? 73.770 67.765 34.057 1.00 30.65 ? ? ? ? ? ? 58 1MA A P 1
ATOM 1253 O OP1 . 1MA A 1 58 ? 72.638 67.886 33.105 1.00 32.84 ? ? ? ? ? ? 58 1MA A OP1 1
ATOM 1254 O OP2 . 1MA A 1 58 ? 73.621 68.229 35.450 1.00 29.49 ? ? ? ? ? ? 58 1MA A OP2 1
ATOM 1255 O "O5'" . 1MA A 1 58 ? 74.315 66.273 34.254 1.00 28.81 ? ? ? ? ? ? 58 1MA A "O5'" 1
ATOM 1256 C "C5'" . 1MA A 1 58 ? 74.592 65.439 33.080 1.00 29.42 ? ? ? ? ? ? 58 1MA A "C5'" 1
ATOM 1257 C "C4'" . 1MA A 1 58 ? 74.279 63.972 33.383 1.00 33.42 ? ? ? ? ? ? 58 1MA A "C4'" 1
ATOM 1258 O "O4'" . 1MA A 1 58 ? 74.880 63.685 34.667 1.00 32.36 ? ? ? ? ? ? 58 1MA A "O4'" 1
ATOM 1259 C "C3'" . 1MA A 1 58 ? 72.789 63.573 33.509 1.00 35.13 ? ? ? ? ? ? 58 1MA A "C3'" 1
ATOM 1260 O "O3'" . 1MA A 1 58 ? 72.625 62.168 33.250 1.00 36.80 ? ? ? ? ? ? 58 1MA A "O3'" 1
ATOM 1261 C "C2'" . 1MA A 1 58 ? 72.560 63.667 35.012 1.00 34.80 ? ? ? ? ? ? 58 1MA A "C2'" 1
ATOM 1262 O "O2'" . 1MA A 1 58 ? 71.525 62.828 35.506 1.00 36.27 ? ? ? ? ? ? 58 1MA A "O2'" 1
ATOM 1263 C "C1'" . 1MA A 1 58 ? 73.908 63.150 35.551 1.00 33.62 ? ? ? ? ? ? 58 1MA A "C1'" 1
ATOM 1264 N N9 . 1MA A 1 58 ? 74.284 63.494 36.930 1.00 30.36 ? ? ? ? ? ? 58 1MA A N9 1
ATOM 1265 C C8 . 1MA A 1 58 ? 73.887 64.574 37.688 1.00 34.55 ? ? ? ? ? ? 58 1MA A C8 1
ATOM 1266 N N7 . 1MA A 1 58 ? 74.415 64.610 38.899 1.00 33.32 ? ? ? ? ? ? 58 1MA A N7 1
ATOM 1267 C C5 . 1MA A 1 58 ? 75.204 63.469 38.953 1.00 33.37 ? ? ? ? ? ? 58 1MA A C5 1
ATOM 1268 C C6 . 1MA A 1 58 ? 76.031 62.941 39.948 1.00 33.58 ? ? ? ? ? ? 58 1MA A C6 1
ATOM 1269 N N6 . 1MA A 1 58 ? 76.184 63.488 41.134 1.00 41.19 ? ? ? ? ? ? 58 1MA A N6 1
ATOM 1270 N N1 . 1MA A 1 58 ? 76.708 61.803 39.669 1.00 34.48 ? ? ? ? ? ? 58 1MA A N1 1
ATOM 1271 C CM1 . 1MA A 1 58 ? 77.649 61.222 40.626 1.00 31.43 ? ? ? ? ? ? 58 1MA A CM1 1
ATOM 1272 C C2 . 1MA A 1 58 ? 76.527 61.216 38.479 1.00 28.43 ? ? ? ? ? ? 58 1MA A C2 1
ATOM 1273 N N3 . 1MA A 1 58 ? 75.793 61.624 37.453 1.00 31.67 ? ? ? ? ? ? 58 1MA A N3 1
ATOM 1274 C C4 . 1MA A 1 58 ? 75.142 62.771 37.747 1.00 33.02 ? ? ? ? ? ? 58 1MA A C4 1
While I have not tested exhaustively, it seems true that PDBx/mmCIF has adopted a different definition of what constitutes a HETATM residue. It is worth noting that results from 3DNA and DSSR/SNAP are not effected by the conflicting assignments.

Nowadays, “big data” and “big science” are hot topics. They all sound good and certainly come about for a reason. Yet, to transform data to information to knowledge to understanding to wisdom, sophisticated software tools are required. The programs can be big and complicated, or small and self-contained, fitting different purposes. As long as they can get the claimed job done in a robust fashion, size should not be a concern.
Over the years, however, I have seen a trend of bloated software with many (fragile) dependencies in bioinformatics. Some tools are so picky and hard to use/maintain that instead of serving, they become sort of a master. As a more representative example, I recently tried to install an open-source software associated with a paper published just a few years ago in a leading journal. The software has only a few dependencies, yet some of them have already become obsolete. I spent hours each time, on Mac OS X and two versions of Ubuntu Linux, but failed to get it running properly (always abort with error messages). The download page hosting the software has been inactive since around the publication of the paper. Presumably, the PhD student or postdoc who wrote the code had left the lab, and with a paper published, all is done!
As an active practitioner of bioinformatics for well over a decade, I can confidently claim to be well above average in familiarity with Linux/Mac OS X and associated shell programming and make etc tools, and various common scripting and compiled programming languages. Yet, once in a while, I get frustrated when I try to download and install a software tool attached to a paper I am interested in. As I see it, the vast majority of software programs from research labs are publication-oriented — as long a paper is published, it is finished.
From my experience, I always see software as engineering. It needs careful design and great attention to meticulous details. A sophisticated piece of scientific software is a combination of science and engineering. Expertise in domain knowledge is a must, and refined skills in computer programming is indispensable. The DSSR program I created and continuously refined over the past three years represents what a scientific software should be in my believe.
Among other unique features, DSSR is tiny (< 1mb), self-contained (without run-time dependencies) and runs on Windows, Mac OS X, and Linux. Getting DSSR up and running should take only minutes by any one with basic familiarity of common computer systems. I have no doubt that the beauty of being small as represented by DSSR will be gradually appreciated by the community.